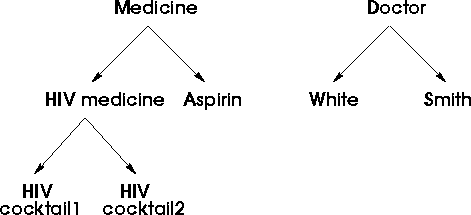
Figure 5: Example of a Taxonomy
We briefly outline our data structure used in our implementation. Typically we will outline how to represent an item or generalized item, how to represent an association rule and how is the counting done with our specially data structure.
We use the following structure to represent a node in the taxonomy tree. Each node represents an item or generalized item if it is not a leaf.
typedef struct node_st
{
struct node_st *parent;
int num_child; /* Number of children, 0 if it is a leaf */
struct node_st *children;
int num_leaves; /* total number of leaves in the subtree */
int attr_num; /* is the corresponding attribute number for
a leaf node, otherwise it is used to hold
the least attribute number of all children
leaves */
int node_num; /* The unique node id */
template bv_template;
int attr_group; /* stores the id of the tree (attribute group)
the corresponding attribute is in */
int count_pos;
int count_neg;
} node;
where is a bit vector, and is defined as:
typedef struct
{
int begin;
int length;
int *bv;
} template;
So is a variable length bit vector.
Consider the following example.

Figure 5: Example of a Taxonomy
In the above taxonomy tree, the template for each node is (assume length 3 of bit vector for medicine tree, and 2 for doctor tree):
Medicine: 111 HIV medicine : 110 HIV cocktail 1: 100 HIV cocktail 2: 010 Aspirin: 001 Doctor: 11 White: 10 Smith: 01
Suppose we have rule . The template for this rule is . A tuple like tuple 1: medicine = HIV cocktail1, doctor = Smith has the following template: , which will match the template of , since the AND of the template and the tuple is , which have bit set to in each attribute node of the rule. So the tuple 1 will support the above template. From this example we can see that we don't need to extend the tuple to count the frequency of the generalized rules, which is done in the IBM algorithms. Also, by using the bit vector, we have a very compact representation for both tuples and candidates, which saves a lot of memeory.